AI in Microscope Image Processing
AI is revolutionizing microscope image processing with powerful capabilities such as precise segmentation, noise reduction, super-resolution, and automated image acquisition. This article highlights essential AI tools and emerging trends in scientific research.
AI techniques are revolutionizing microscopy by optimizing image acquisition and automating analysis. In modern smart microscopes, AI modules can adjust imaging parameters on-the-fly (e.g., focus, illumination) to minimize photobleaching and enhance signal. Meanwhile, deep learning algorithms can sift through complex image data to extract hidden biological insights and even link images to other data (e.g., genomics).
AI Methods: Machine Learning vs Deep Learning
AI methods range from classic machine learning (ML) to modern deep learning (DL). Each approach has distinct strengths and limitations:
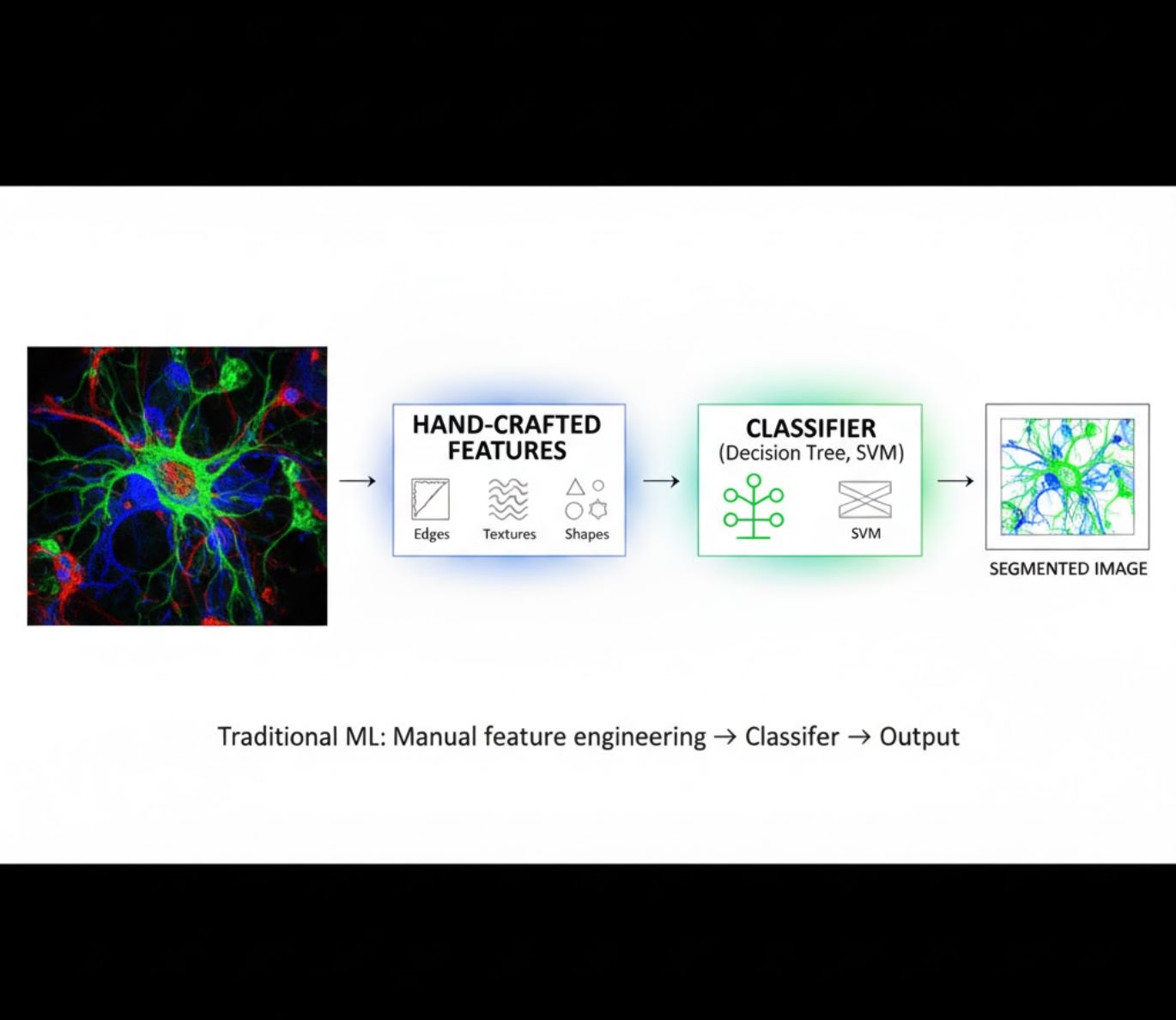
Hand-Crafted Features
- Researchers manually craft image features (edges, textures, shapes)
- Features fed to classifiers (decision trees, SVM)
- Fast to train
- Struggles with complex or noisy images
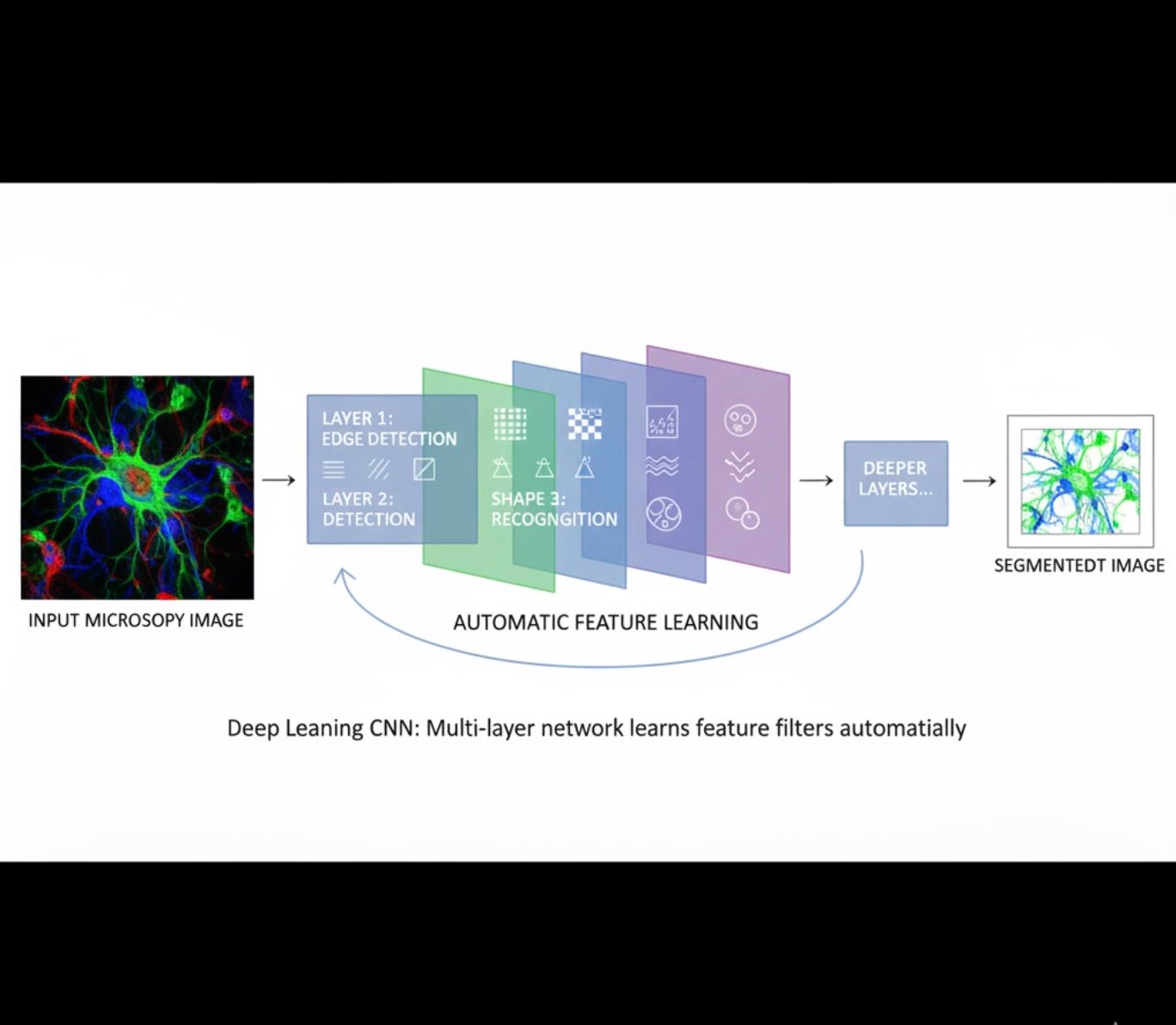
Automatic Feature Learning
- Multi-layer neural networks (CNNs) learn features automatically
- End-to-end learning from raw pixels
- Much more robust to variations
- Captures intricate textures and structures reliably
How CNNs work: Convolutional neural networks apply successive filters to microscopy images, learning to detect simple patterns (edges) in early layers and complex structures (cell shapes, textures) in deeper layers. This hierarchical learning makes DL exceptionally robust even when intensity profiles vary significantly.
Visual Comparison: ML vs DL Pipelines
Key AI Applications in Microscopy
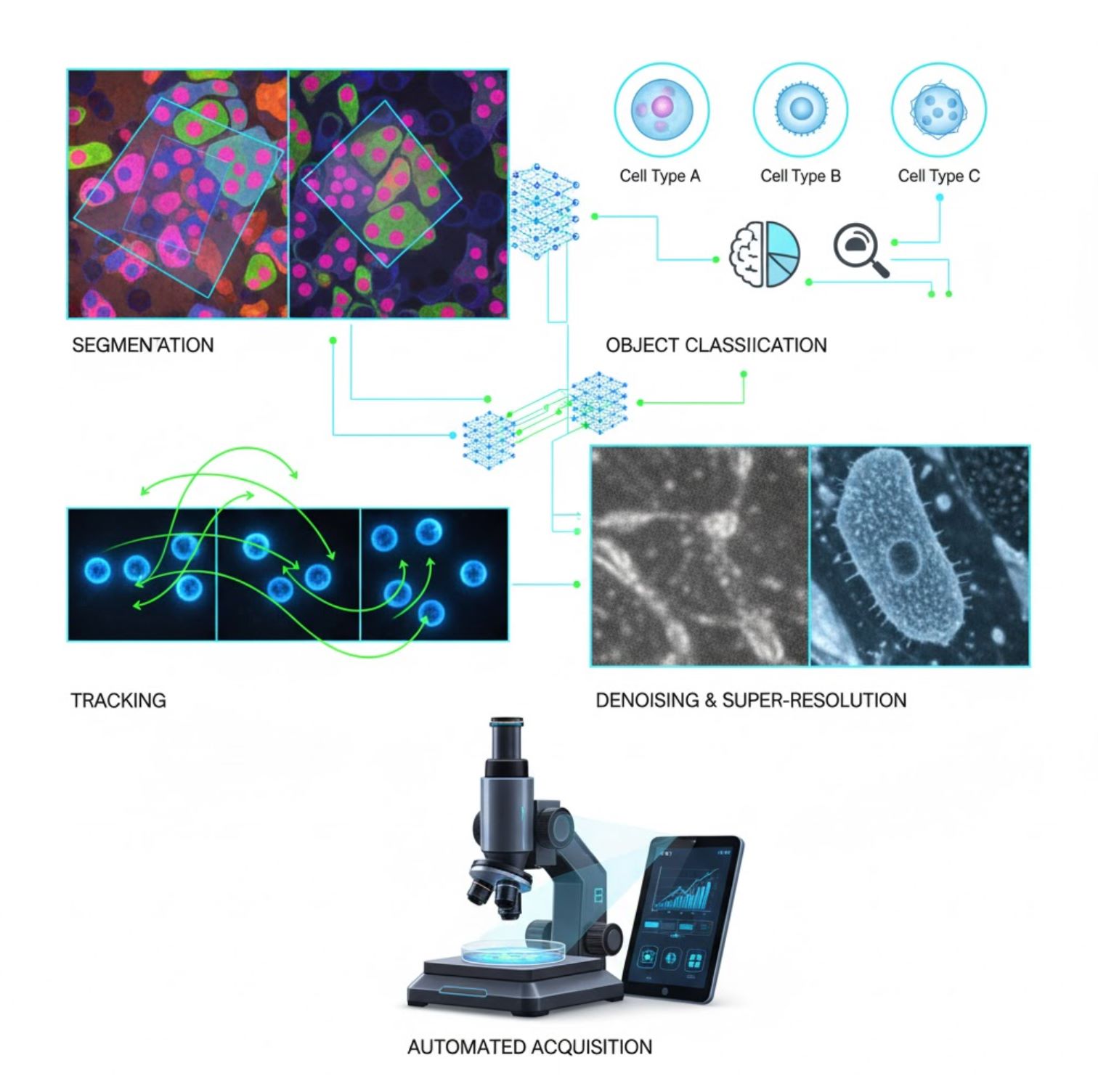
AI is now embedded in many image-processing tasks across the microscopy workflow:
Segmentation
Partitioning images into regions (e.g., identifying every cell or nucleus). Deep networks like U-Net excel at this task.
- Semantic segmentation: Per-pixel class labels
- Instance segmentation: Separating individual objects
- High accuracy on crowded or dim images
- Vision foundation models (e.g., μSAM) now adapted for microscopy
Object Classification
After segmentation, AI classifies each object with high precision.
- Cell type identification
- Mitotic stage determination
- Pathology indicator detection
- Distinguishes subtle phenotypes hard to quantify manually
Tracking
In time-lapse microscopy, AI tracks cells or particles across frames with unprecedented accuracy.
- Deep learning dramatically improves tracking accuracy
- Enables reliable analysis of moving cells
- Captures dynamic biological processes
Denoising & Super-Resolution
AI models enhance image quality by removing noise and blur.
- Physics-informed deep models learn microscope optics
- Reconstruct sharper, artifact-free images
- Higher resolution with reduced artifacts vs. traditional methods
Automated Acquisition
AI guides the microscope itself in real-time.
- Analyzes live images to make intelligent decisions
- Automatically adjusts focus and scans areas of interest
- Reduces phototoxicity and saves time
- Enables high-throughput and adaptive imaging experiments
Popular AI Tools in Microscope Image Processing
A rich ecosystem of tools supports AI in microscopy. Researchers have built both general-purpose and specialized software, many open-source:
Cellpose
| Developer | Carsen Stringer and Marius Pachitariu (MouseLand research group) |
| Supported Platforms |
Requires Python (pip/conda installation). GUI available on desktop only. |
| Language Support | English documentation; globally adopted in research labs worldwide |
| Pricing Model | Free and open-source under BSD-3-Clause license |
Overview
Cellpose is an advanced, deep-learning–based segmentation tool designed for microscopy images. As a generalist algorithm, it accurately segments diverse cell types (nuclei, cytoplasm, etc.) across different imaging modalities without requiring model retraining. With human-in-the-loop capabilities, researchers can refine results, adapt the model to their data, and apply the system to both 2D and 3D imaging workflows.
Key Features
Works out of the box for a wide variety of cell types, stains, and imaging modalities without custom training.
Supports full 3D stacks using a "2.5D" approach that reuses 2D models for volumetric data.
Manually correct segmentation results and retrain the model on your custom data for improved accuracy.
Access via Python API, command-line interface, or graphical user interface for flexible workflows.
Denoising, deblurring, and upsampling capabilities to enhance image quality before segmentation.
Download or Access
Technical Background
Cellpose was introduced in a seminal study by Stringer, Wang, Michaelos, and Pachitariu, trained on a large and highly varied dataset containing over 70,000 segmented objects. This diversity enables the model to generalize across cell shapes, sizes, and microscopy settings, significantly reducing the need for custom training in most use cases. For 3D data, Cellpose cleverly reuses its 2D model in a "2.5D" fashion, avoiding the need for fully 3D-annotated training data while still delivering volumetric segmentation. Cellpose 2.0 introduced human-in-the-loop retraining, allowing users to manually correct predictions and retrain on their own images for improved performance on specific datasets.
Installation & Setup
Set up a Python environment using conda:
conda create -n cellpose python=3.10Activate the environment and install Cellpose:
# For GUI support
pip install cellpose[gui]
# For minimal setup (API/CLI only)
pip install cellposeGetting Started
GUI Mode
- Launch the GUI by running:
python -m cellpose - Drag and drop image files (
.tif,.png, etc.) into the interface - Select model type (e.g., "cyto" for cytoplasm or "nuclei" for nuclei)
- Set estimated cell diameter or let Cellpose auto-calibrate
- Click to start segmentation and view results
Python API Mode
from cellpose import models
# Load model
model = models.Cellpose(model_type='cyto')
# Segment images
masks, flows = model.eval(images, diameter=30)Refine & Retrain
- After generating masks, correct segmentation in the GUI by merging or deleting masks manually
- Use built-in training functions to retrain on corrected examples
- Improved model performance on your specific dataset
Process 3D Data
- Load a multi-Z TIFF or volumetric stack
- Use the
--Zstackflag in GUI or API to process as 3D - Optionally refine 3D flows via smoothing or specialized parameters for better segmentation
Limitations & Considerations
- Model Generality Trade-off: While the generalist model works broadly, highly unusual cell shapes or imaging conditions may require retraining.
- Annotation Effort: Human-in-the-loop training requires manual corrections, which can be time-consuming for large datasets.
- Installation Complexity: GUI installation may require command-line use, conda environments, and managing Python dependencies — not always straightforward for non-programmers.
- Desktop Only: Cellpose is designed for desktop use; no native Android or iOS applications available.
Frequently Asked Questions
No — Cellpose provides pre-trained, generalist models that often work well without retraining. However, for optimal results on special or unusual data, you can annotate and retrain using the human-in-the-loop features.
Yes — it supports 3D by reusing its 2D model (so-called "2.5D"), and you can run volumetric stacks through the GUI or API.
A GPU is highly recommended for faster inference and training, especially on large or 3D datasets, but Cellpose can run on CPU-only machines with slower performance.
In the GUI, set the estimated cell diameter manually or let Cellpose automatically calibrate it. You can refine results and retrain if segmentation is not optimal.
Yes — newer versions (Cellpose 3) include image restoration models for denoising, deblurring, and upsampling to improve segmentation quality before processing.
StarDist
| Developer | Uwe Schmidt, Martin Weigert, Coleman Broaddus, and Gene Myers |
| Supported Platforms |
|
| Language Support | Open-source project with documentation and community primarily in English |
| Pricing Model | Free and open source. Licensed under BSD-3-Clause |
Overview
StarDist is a deep-learning tool for instance segmentation in microscopy images. It represents each object (such as cell nuclei) as a star-convex polygon in 2D or polyhedron in 3D, enabling accurate detection and separation of densely packed or overlapping objects. With its robust architecture, StarDist is widely used for automated cell and nucleus segmentation in fluorescence microscopy, histopathology, and other bioimage analysis applications.
Key Features
Highly accurate instance segmentation using star-convex polygons (2D) and polyhedra (3D) for reliable object detection.
Dedicated models for both 2D images and 3D volumetric data for comprehensive microscopy analysis.
Ready-to-use models for fluorescence nuclei, H&E-stained histology, and other common imaging scenarios.
Classify detected objects into distinct classes (e.g., different cell types) in a single segmentation run.
Seamless integration with ImageJ/Fiji, QuPath, and napari for accessible GUI-based workflows.
Comprehensive instance segmentation evaluation including precision, recall, F1 score, and panoptic quality.
Technical Background
Originally introduced in a MICCAI 2018 paper, StarDist's core innovation is the prediction of radial distances along fixed rays combined with object probability for each pixel, enabling accurate reconstruction of star-convex shapes. This approach reliably segments closely touching objects that are difficult to separate using traditional pixel-based or bounding-box methods.
Recent developments have expanded StarDist to histopathology images, enabling not only nucleus segmentation but also multi-class classification of detected objects. The method achieved top performance in challenges such as the CoNIC (Colon Nuclei Identification and Counting) challenge.
Download or Access
Installation & Setup
Install TensorFlow (version 1.x or 2.x) as a prerequisite for StarDist.
Use pip to install the StarDist Python package:
pip install stardistFor napari:
pip install stardist-napariFor QuPath: Install the StarDist extension by dragging the .jar file into QuPath.
For ImageJ/Fiji: Use the built-in plugin manager or manual installation via the plugins menu.
Running Segmentation
Load a pre-trained model, normalize your image, and run prediction:
from stardist.models import StarDist2D
model = StarDist2D.from_pretrained('2D_versatile_fluo')
labels, details = model.predict_instances(image)Open your image in napari, select the StarDist plugin, choose a pre-trained or custom model, and run prediction directly from the GUI.
Use the StarDist plugin from the Plugins menu to apply a model on your image stack with an intuitive interface.
After installing the extension, run StarDist detection via QuPath's scripting console or graphical interface for histopathology analysis.
Training & Fine-Tuning
Create ground-truth label images where each object is uniquely labeled. Use annotation tools like LabKit, QuPath, or Fiji to prepare your dataset.
Use StarDist's Python API to train a new model or fine-tune an existing one with your custom annotated data.
Post-Processing Options
- Apply non-maximum suppression (NMS) to eliminate redundant candidate shapes
- Use StarDist OPP (Object Post-Processing) to merge masks for non-star-convex shapes
Limitations & Considerations
- Star-convex assumption may not model highly non-convex or very irregular shapes perfectly
- Installation complexity: custom installs require a compatible C++ compiler for building extensions
- GPU acceleration depends on compatible TensorFlow, CUDA, and cuDNN versions
- Some users report issues running the ImageJ plugin due to Java configuration
Frequently Asked Questions
StarDist works with a variety of image types including fluorescence, brightfield, and histopathology (e.g., H&E), thanks to its flexible pre-trained models and adaptability to different imaging modalities.
Yes — StarDist supports 3D instance segmentation using star-convex polyhedra for volumetric data, extending the 2D capabilities to full 3D analysis.
Not necessarily. Pre-trained models are available and often work well out-of-the-box. However, for specialized or novel data, annotating and training custom models improves accuracy significantly.
StarDist integrates with napari, ImageJ/Fiji, and QuPath, allowing you to run segmentation from a GUI without coding. It also supports direct Python API usage for advanced workflows.
StarDist provides built-in functions for computing common instance segmentation metrics including precision, recall, F1 score, and panoptic quality to assess segmentation performance.
SAM
Application Information
| Developer | Meta AI Research (FAIR) |
| Supported Devices |
|
| Language & Availability | Open-source foundation model available globally; documentation in English |
| Pricing | Free — open-source under Meta's license via GitHub and MIB integration |
General Overview
SAM (Segment Anything Model) is a powerful AI foundation model created by Meta that enables interactive and automatic segmentation of virtually any object in images. Using prompts such as points, bounding boxes, or rough masks, SAM generates segmentation masks without requiring task-specific retraining. In microscopy research, SAM's flexibility has been adapted for cell segmentation, organelle detection, and histopathology analysis, offering a scalable solution for researchers needing a promptable, general-purpose segmentation tool.
Detailed Introduction
Originally trained by Meta on over 1 billion masks across 11 million images, SAM was designed as a promptable foundation model for segmentation with "zero-shot" performance on novel domains. In medical imaging research, SAM has been evaluated for whole-slide pathology segmentation, tumor detection, and cell nuclei identification. However, its performance on densely packed instances—such as cell nuclei—is mixed: even with extensive prompts (e.g., 20 clicks or boxes), zero-shot segmentation can struggle in complex microscopy images.
To address this limitation, domain-specific adaptations have emerged:
- SAMCell — Fine-tuned on large microscopy datasets for strong zero-shot segmentation across diverse cell types without per-experiment retraining
- μSAM — Retrained on over 17,000 manually annotated microscopy images to improve accuracy on small cellular structures
Key Features
Flexible interaction using points, boxes, and masks for precise control.
Performs segmentation without fine-tuning on new image domains.
Adaptable for microscopy and histopathology via few-shot or prompt-based retraining.
Available in Microscopy Image Browser (MIB) with 3D and interpolated segmentation support.
IDCC-SAM enables automatic cell counting in immunocytochemistry without manual annotation.
Download or Access
User Guide
- Open Microscopy Image Browser and navigate to the SAM segmentation panel
- Configure the Python interpreter and select between SAM-1 or SAM-2 models
- For GPU acceleration, select "cuda" in the execution environment (recommended for optimal performance)
- Point prompts: Click on an object to define a positive seed; use Shift + click to expand and Ctrl + click for negative seeds
- 3D stacks: Use Interactive 3D mode—click on one slice, shift-scroll, and interpolate seeds across slices
- Adjust mode: Replace, add, subtract masks, or create a new layer as needed
- Use MIB's "Automatic everything" option in the SAM-2 panel to segment all visible objects in a region
- Review and refine masks after segmentation as needed
- Use prompt-based fine-tuning pipelines (e.g., "All-in-SAM") to generate pixel-level annotations from sparse user prompts
- For cell counting, apply IDCC-SAM, which uses SAM in a zero-shot pipeline with post-processing
- For high-accuracy cell segmentation, use SAMCell, fine-tuned on microscopy cell images
Limitations & Considerations
- Zero-shot performance inconsistent on dense or overlapping structures without domain tuning
- Segmentation quality depends heavily on prompt design and strategy
- GPU strongly recommended; CPU inference is very slow
- Struggles with very high-resolution whole-slide images and multi-scale tissue structures
- Fine-tuning or adapting SAM for microscopy may require machine learning proficiency
Frequently Asked Questions
Yes—through adaptations like SAMCell, which fine-tunes SAM on microscopy datasets specifically for cell segmentation tasks.
Not always. With IDCC-SAM, you can perform zero-shot cell counting without manual annotations.
Use prompt-based fine-tuning (e.g., "All-in-SAM") or pretrained microscopy versions like μSAM, which is trained on over 17,000 annotated microscopy images.
While possible on CPU, GPU is highly recommended for practical inference speed and real-time interactive segmentation.
Yes—MIB's SAM-2 integration supports 3D segmentation with seed interpolation across slices for volumetric analysis.
AxonDeepSeg
| Developer | NeuroPoly Lab at Polytechnique Montréal and Université de Montréal |
| Supported Platforms |
|
| Language | English documentation; open-source tool used globally |
| Pricing | Free and open-source |
Overview
AxonDeepSeg is an AI-powered tool for automatic segmentation of axons and myelin in microscopy images. Using convolutional neural networks, it delivers accurate three-class segmentation (axon, myelin, background) across multiple imaging modalities including TEM, SEM, and bright-field microscopy. By automating morphometric measurements such as axon diameter, g-ratio, and myelin thickness, AxonDeepSeg streamlines quantitative analysis in neuroscience research, significantly reducing manual annotation time and improving reproducibility.
Key Features
Ready-to-use models optimized for TEM, SEM, and bright-field microscopy modalities.
Precise classification of axon, myelin, and background regions in microscopy images.
Automatic computation of axon diameter, g-ratio, myelin thickness, and density metrics.
Napari GUI integration enables manual refinement of segmentation masks for enhanced accuracy.
Integrates seamlessly into custom pipelines for large-scale neural tissue analysis.
Comprehensive test scripts ensure reproducibility and reliable segmentation results.
Technical Details
Developed by the NeuroPoly Lab, AxonDeepSeg leverages deep learning to deliver high-precision segmentation for neuroscientific applications. Pre-trained models are available for different microscopy modalities, ensuring versatility across imaging techniques. The tool integrates with Napari, allowing interactive corrections of segmentation masks, which enhances accuracy on challenging datasets. AxonDeepSeg computes key morphometric metrics, supporting high-throughput studies of neural tissue structure and pathology. Its Python-based framework enables integration into custom pipelines for large-scale analysis of axon and myelin morphology.
Download or Access
Installation & Setup
Ensure Python 3.8 or later is installed, then install AxonDeepSeg and Napari using pip:
pip install axondeepseg napariRun the provided test scripts to confirm all components are properly installed and functioning.
Import microscopy images (TEM, SEM, or bright-field) into Napari or your Python environment.
Choose the appropriate pre-trained model for your imaging modality and run segmentation to generate axon and myelin masks.
Automatically compute morphometric measurements including axon diameter, g-ratio, density, and myelin thickness, then export results in CSV format.
Use the Napari GUI to manually adjust segmentation masks where needed, merging or deleting masks for improved accuracy.
Important Considerations
- Performance may decrease on novel or untrained imaging modalities
- Manual corrections may be required for challenging or complex regions
- GPU is recommended for faster processing of large datasets; CPU processing is also supported
Frequently Asked Questions
AxonDeepSeg supports TEM (Transmission Electron Microscopy), SEM (Scanning Electron Microscopy), and bright-field microscopy with pre-trained models optimized for each modality.
Yes, AxonDeepSeg is completely free and open-source, available for academic and commercial use.
Yes, AxonDeepSeg automatically calculates axon diameter, g-ratio, myelin thickness, and density metrics from segmented images.
GPU is recommended for faster segmentation of large datasets, but CPU processing is also supported for smaller analyses.
Yes, Napari GUI integration allows interactive corrections and refinement of segmentation masks for higher accuracy on challenging regions.
Ilastik
| Developer | Ilastik Team at the European Molecular Biology Laboratory (EMBL) and associated academic partners |
| Supported Platforms |
|
| Language | English |
| Pricing | Free and open-source |
Overview
Ilastik is a powerful, AI-driven tool for interactive image segmentation, classification, and analysis of microscopy data. Using machine learning techniques like Random Forest classifiers, it enables researchers to segment pixels, classify objects, track cells over time, and perform density counting in both 2D and 3D datasets. With its intuitive interface and real-time feedback, Ilastik is accessible to scientists without programming expertise and is widely adopted in cell biology, neuroscience, and biomedical imaging.
Key Features
Real-time feedback as you annotate representative regions for instant segmentation results.
Categorize segmented structures based on morphological and intensity features.
Track cell movement and division in 2D and 3D time-lapse microscopy experiments.
Quantify crowded regions without explicit segmentation of individual objects.
Semi-automatic segmentation for complex 3D volumes with intuitive interaction.
Process multiple images automatically using headless command-line mode.
Download
Getting Started Guide
Download Ilastik for your operating system from the official website. The package includes all necessary Python dependencies, so follow the installation instructions for your platform.
Open Ilastik and choose your analysis workflow: Pixel Classification, Object Classification, Tracking, or Density Counting. Load your image dataset, which can include multi-channel, 3D, or time-lapse images.
Label a few representative pixels or objects in your images. Ilastik's Random Forest classifier learns from these annotations and automatically predicts labels across your entire dataset.
Apply the trained model to segment or classify your full dataset. Export results as labeled images, probability maps, or quantitative tables for downstream analysis and visualization.
Use Ilastik's headless mode to automatically process multiple images without manual intervention, ideal for large-scale analysis pipelines.
Limitations & Considerations
- Interactive labeling can be time-consuming for very large datasets
- Accuracy depends on the quality and representativeness of user annotations
- Memory requirements — very high-resolution or multi-gigabyte datasets may require significant RAM
- Complex data — Random Forest classifiers may underperform compared to deep neural networks on highly variable or complex imaging data
Frequently Asked Questions
Yes, Ilastik fully supports 3D volumes and time-lapse experiments for segmentation, tracking, and quantitative analysis across multiple timepoints.
Yes, Ilastik is completely free and open-source, available for all users without licensing restrictions.
No, Ilastik provides an intuitive graphical interface with real-time feedback, making it accessible to researchers without programming expertise. Advanced users can also use command-line batch processing.
Yes, the dedicated tracking workflow enables analysis of cell movement and division in both 2D and 3D time-lapse datasets with automatic lineage tracking.
Segmentation outputs can be exported as labeled images, probability maps, or quantitative tables, allowing seamless integration with downstream analysis tools and visualization software.
These tools span novice to expert levels. Many are free and open-source, facilitating reproducible and shareable AI workflows across the research community.
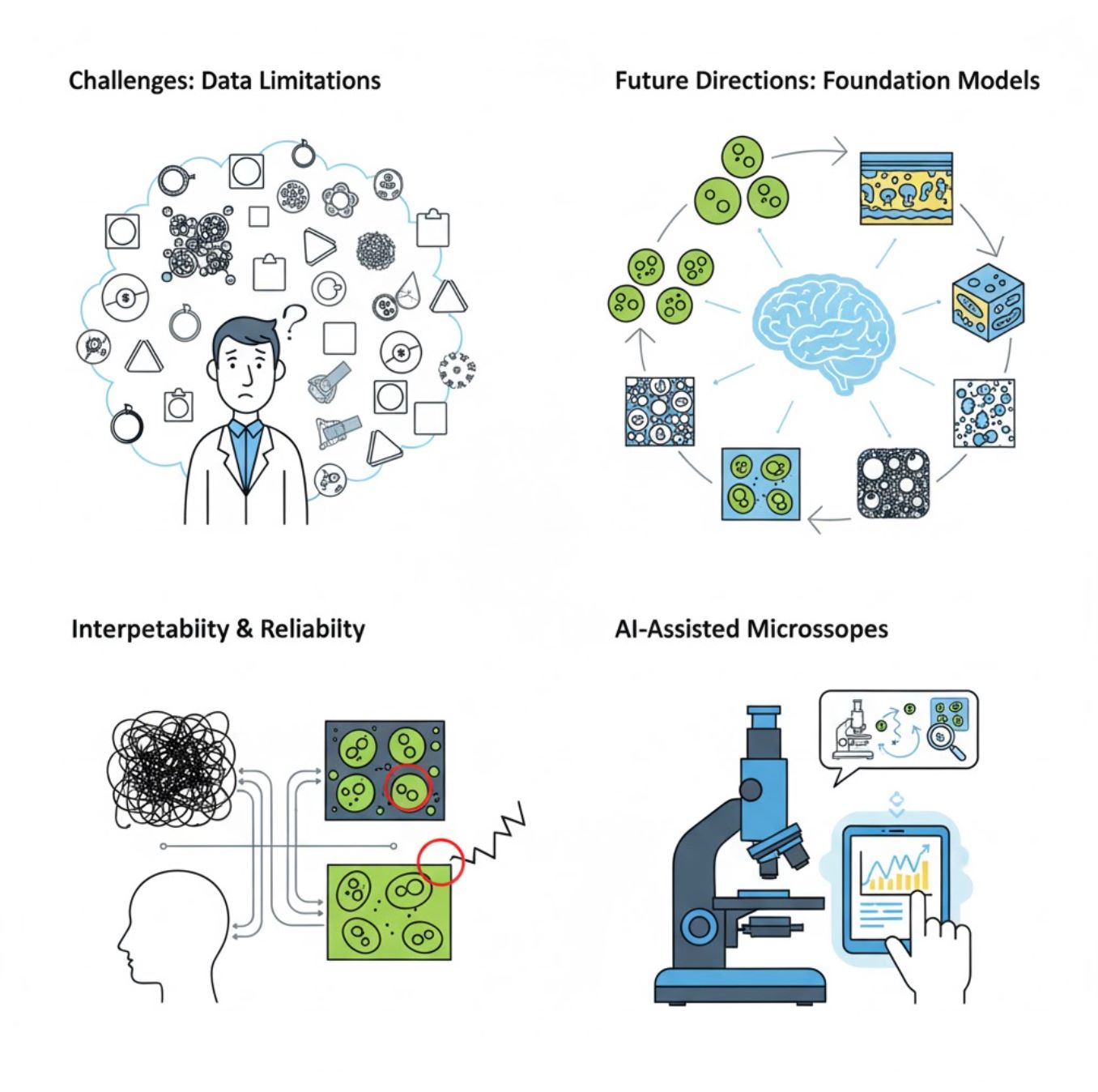
Challenges and Future Directions
Current Challenges
Emerging Trends
Vision Foundation Models
Next-generation AI systems promise to reduce the need for task-specific training.
- Models like SAM and CLIP-based systems
- One AI handles many microscopy tasks
- Faster deployment and adaptation
AI-Assisted Microscopes
Fully autonomous and intelligent microscopy systems are becoming reality.
- Natural language control via LLMs
- Fully automated feedback loops
- Democratizes advanced microscopy access
Key Takeaways
- AI is rapidly transforming microscope image processing with improved accuracy and automation
- Deep learning outperforms traditional machine learning on complex, variable microscopy images
- CNNs automatically learn hierarchical features from raw pixels for robust analysis
- Key applications include segmentation, classification, tracking, denoising, and automated acquisition
- Success depends on quality data and careful validation by experts
- Vision foundation models and AI-assisted microscopes represent the future of the field
With continued advances and community efforts (open-source tools, shared datasets), AI will increasingly become a core part of the microscope's "eye," helping scientists see the unseen.







No comments yet. Be the first to comment!